K-CLIER: A clinical application of single-cell RNA sequencing data
In collaboration with Dr. Med. Pietro Cippà, Dr. Anna Rinaldi and Lorenzo Ruinelli (EOC)
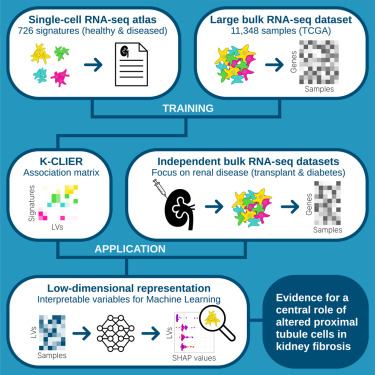
Single-cell transcriptomics is an extremely rich source of data, which has been crucial in refining our understanding of the cellular composition of tissues and organs. However, unlike bulk transcriptomics, the use of such technology in clinical practice faces major challenges related to tissue processing and data analysis.
With K-CLIER (Legouis, Rinaldi, Malpetti, et al., 2024), we used state-of-the-art transfer learning methods to learn a transformation that produces a low-dimensional representation of bulk gene expression that can be easily interpreted from a single-cell perspective.
We applied this transformation to various datasets from patients with renal pathologies. Using the transformed data, we developed machine learning models to predict clinical outcomes and therapy response in oncological patients, and we studied rejection in transplanted patients. Notably, this approach could be extended to other medical contexts beyond renal applications (Malpetti, Mangili, et al., 2025).